3D Chromosome Viewer
Contents |
Background and Project Overview
Most genome browsers display DNA linearly, using single-dimensional depictions that are useful to examine certain epigenetic mechanisms such as DNA methylation. However, these representations are insufficient to visualize intra-chromosomal interactions and relationships between distal genome features. Relationships between DNA regions may be difficult to decipher or missed entirely if those regions are distant in one dimension but could be spatially proximal when mapped to three-dimensional space. For example, the visualization of enhancers folding over genes is only fully expressed in three-dimensional space. Thus, to accurately understand DNA behavior during gene expression, a means to model chromosomes is essential. The purpose of this project is to facilitate three-dimensional chromosome modelling.
The three-dimensional chromosome model will contain a three-tier model of visualization where each subsequent tier is a greater detailed segment of the preceding tier. At the highest level, the first tier models entire chromosomes consisting of interconnected cylinders. Each cylinder in the first tier represents a chromosomal domain that links to the second tier of visualization. At the next level, the second tier models chromosome domains consisting of cylinders representing bins of 20,000 and 40,000 base pairs. The second tier also depicts genomic features--such as genes, enhancers, and CTCF binding sites-- with different color schemes that elucidate interactions between regions. Each cylinder in the second tier links to the third and final tier, which is a genome browser displaying a detailed linear annotation of the corresponding bin. As a stretch goal, the third tier could also render a helix structure of color-coded base pairs.
The potential applications of this project include the visualization of intra-chromosomal interactions, genomic features and their relationships, and the discovery of new genes.
Screenshots
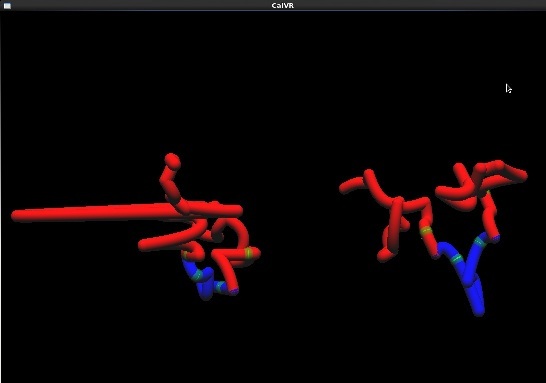
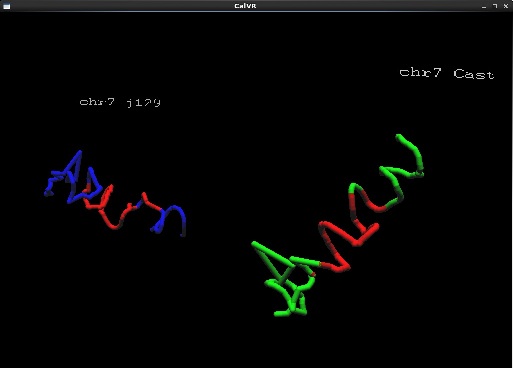
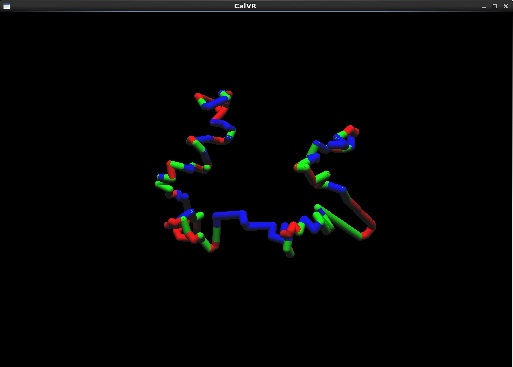
- A few captured screenshots captured on the desktop version of CalVR. Models are rendered at low polygon and tessellation count to account for desktop version.
Status
- Loaded world and menu for initial set up of view
- Added functionality to parse data files to extract data needed for rendering
- Rendered spheres at x,y,z coordinates specified in data files
- Created mathematically manipulated cylinders to connect spheres to create a contiguous structure
- Added options to render any of 23 chromosomes from the menu
- Added colour to differentiate between loose, condensed, and intermediate areas of the chromosome
- Added clear all button
- Switched to using SceneObjects to allow multiple chromosomes to be rendered on the same scene
- Added new datapoints for mouse Chr7 (two versions: mother and father)
- Added new datapoints for cast and j129 ch_3 domains
- Switched to using capsules for optimized rendering (1 drawable instead of 3)
- Switched to using backface culling for optimized rendering
- Using bezier curve matlab script to generate more data points from a set of datapoints for more detailed curve
- Added homologous chromosome data support with colors based on eigen values generated from our sequencing scripts
- Rendered domain models that show influence of CTCF sites
To-Do
- Implement transition to second tier functionality
- Display domain of chromosome bin on click
- Add animation
- Add colour scheme to differentiate different points of interest on specific chromosomes
- Add labels
- Auto adjust camera upon loading of new chromosome
Future Work
- Better interactivity between the next two tiers of the three-tier model
- Add functionality for scripting support
- Add support for rendering description files dynamically
- Write honors thesis/research paper
Possible Extensions
- Is it practical to extend to mobile applications?
- Mobile version of website
- Mobile application
- Mobile experience of VR and other scientific applications using VR
Participants
Software Developers:
Project Advisors:
Misc. Development Assistance:
- Philip Weber
- Andrew Prudhomme
- Alfred Tarng